Protein Packing
From Info
Contents |
Introduction
Changes in protein conformation play key roles in facilitating various biochemical processes, ranging from signaling and phosphorylation to transport and catalysis. While various factors that drive these motions such as environmental changes and binding of small molecules are well understood, specific causative effects on the structural features of the protein due to these conformational changes have not been studied on a large scale.
Methods and Results
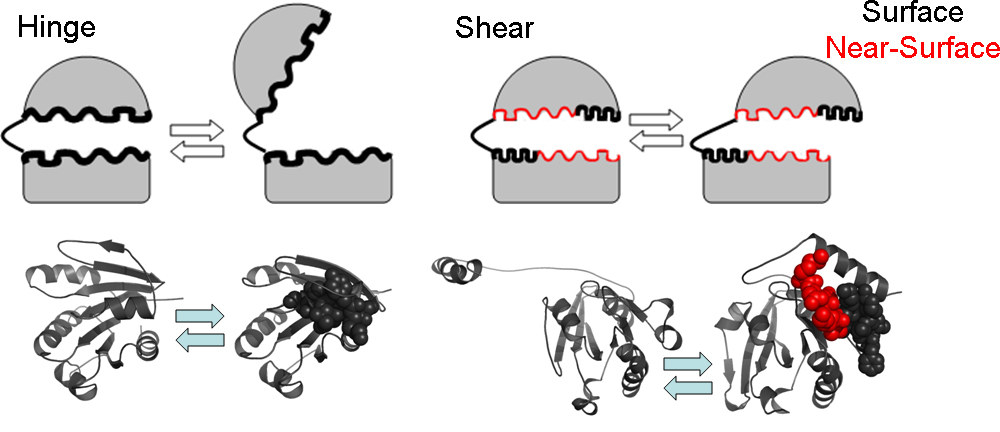
Here, we study protein conformational changes in relation to two key structural metrics: packing efficiency and disorder. Packing has been shown to be crucial for protein stability and function by many protein design and engineering studies. We study changes in packing efficiency during conformational changes, thus extending the analysis from a static context to a dynamic perspective and report some interesting observations. First, we study various proteins that adopt alternate conformations and find that tendencies to show motion and change in packing efficiency are correlated: residues that change their packing efficiency show larger motions. Second, our results suggest that residues that show higher changes in packing during motion are located on the changing interfaces which are formed during these conformational changes. These changing interfaces are slightly different from the shear or static interfaces that have been analyzed in previous studies. Third, analysis of packing efficiency changes in the context of secondary structure shows that, as expected, residues buried in helices show the least change in packing efficiency whereas those embedded in bends are most likely to change packing. Finally, by relating protein disorder to motions, we show that marginally disordered residues which are ordered enough to be crystallized but have sequence patterns indicative of disorder show higher dislocation and a higher change in packing than ordered ones and are located mostly on the changing interfaces.
Conclusion
Overall, our results demonstrate that between the two conformations, the cores of the proteins remain mostly intact while the interfaces display the most elasticity, both in terms of disorder and change in packing efficiency. By doing a variety of tests, we also show that our observations are robust to the solvation state of the proteins.
Supplementary data:
[1] Master Sheet. This sheet can be rearranged to mine the data easily: eg., to get the list of residues that show motion, change their PE, change their exposed surface area.
[2] Readme file for the master sheet explaining each column.
[3] Protein Set used with different kinds of motions.
[4] Other Supplementary Materials (Text and Figures)